Association of advanced glycation end products in Dupuytren disease | Journal of Orthopaedic Surgery and Research
Patients and preparation
The protocol of this study was approved by the institutional review board of Kobe University Hospital. Normal palmar fasciae from five patients with carpal tunnel syndrome (CTS) (control group) and Dupuytren’s morbid cords from five patients without diabetes mellitus (DD group) were harvested during the surgical treatment. We confirmed that both CTS and DD patients did not suffer from diabetes through a pre-operative blood examination. The tissues were frozen quickly at − 80 °C, and a part of the tissues was prepared for biochemical measurements. The remaining tissues were cut into 10-μm-thick sections for staining. The slides were dried at room temperature and fixed in 4% paraformaldehyde.
For in vitro analysis, part of the excised tissue samples was also cut into small pieces (1–2 mm) and plated onto dishes with Dulbecco’s Modified Eagle’s Medium (Sigma, St. Louis, MO, USA) supplemented with 10% fetal bovine serum) (Sigma, St. Louis, MO, USA) and 1% penicillin–streptomycin (Sigma, St. Louis, MO, USA) (regular medium) under sterile conditions. The dishes were maintained in a 5% CO2 chamber at 37 °C. After adhesion of cells to the dishes was observed, the cells were washed in phosphate-buffered saline, detached by using 0.05% trypsin–0.02% EDTA (Wako, Osaka, Japan), and pelleted in 75-cm2 cell culture flasks with regular medium. The cells were then seeded onto 12-well plates at a concentration of 5.0 × 104 cells per well. After the cells had reached 70% confluence, they were cultured in four groups: palmar fascia-derived cells from carpal tunnel syndrome (CTS) patients with regular medium only (control group), palmar fascia-derived cells from CTS patients with regular medium plus AGEs (200 μg/ml, Trans Genic, Kobe, Japan) (control + AGEs group), palmar fascia-derived cells from DD patients with regular medium only (DD group), and palmar fascia-derived cells from DD patients with regular medium plus AGEs (DD + AGEs group). All experiments were performed with cells from the second or third passage, and the same passage of cells was used for each experiment.
Hematoxylin-eosin (HE) staining and immunohistochemistry
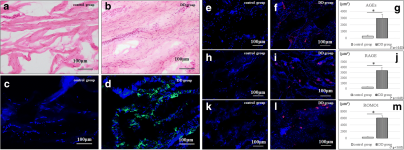
HE staining was performed to observe the histological differences between the control group and DD group. The sections were incubated with alpha-smooth muscle actin (alpha-SMA) antibody (Sigma-Aldrich, St Louis, MO) to confirm the expression of myofibroblasts that characterize DD. The sections were also incubated at 4 °C overnight with the following primary antibodies: anti-AGEs antibody (10 μg/ml; Abcam, Cambridge, UK), anti-RAGE antibody (10 μg/ml; Abcam), and anti-reactive oxygen species modulator 1 (ROMO1) antibody (0.7 μg/ml; Abcam). After incubation with the primary antibodies, the sections were incubated with the secondary antibody, Alexa Fluor 594 (20 μg/ml; Abcam), and counterstained with 4′,6-diamidino-2-phenylindole (DAPI). Digital images of AGEs, RAGE, ROMO1 (red), and nuclear (blue) staining were captured by using a Bio Zero BZ-8000 (Keyence, Osaka, Japan). The positive areas were quantified by using selective coloring in Adobe Photoshop (CS6; Adobe Systems, San Jose, CA, USA) [13].
Expression of superoxide in the cells
After 3 days of exposure to four types of media (control group, control + AGEs group, DD group, DD + AGEs group), superoxide detection reagent was detected by using a Total ROS/Superoxide detection kit (Enzo Life Science, Plymouth Meeting, PA, USA), and the cells were counterstained with DAPI. Digital images were captured from five non-overlapping microscopic fields per well by using the Bio Zero BZ-8000 (Keyence, Osaka, Japan), and superoxide-positive cells were counted.
Real-time polymerase chain reaction (PCR)
Total RNA was extracted from the cells and the tissues by using an RNeasy Mini Kit (Qiagen, Valencia, CA, USA). Total RNA was reverse transcribed into single-strand cDNA by using a High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA). A real-time PCR was performed in triplicate on the cDNA with an Applied Biosystems 7900HT Fast Real-time PCR System and SYBR Green reagents (Applied Biosystems, Foster City, CA, USA). The results were normalized to housekeeping gene expression levels and expressed relative to the control (untreated) culture levels by using the 2-∆∆Ct method. We used the primers for RAGE, nicotinamide adenine dinucleotide phosphate oxidase (Nox)-1, and Nox-4.
The following primer sequences were used: GAPDH: F 5′-CCA CCC ATG GCA AAT TCC-3′ R 5′-GAT GGG ATT TCC ATT GAT GAC A-3′; RAGE: F 5′-AGG AGC GTG CAG AAC TGA AT-3′ R 5′-TTG GCA AGG TGG GGT TAT AC-3′; Nox1: F 5′-GTA CAA ATT CCA GTG TGC AGA CCA C-3′ R 5′-CAG ACT GGA ATA TCG GTG ACA GCA-3′; Nox4: F 5′-CTC AGC GGA ATC AAT CAG CTG TG-3′ R 5′-AGA GGA ACA CGA CAA TCA GCC TTA G-3′.
Statistical analysis
Data are presented as means ± standard deviations of independent experiments. These analyses were performed by using EZR (Saitama Medical Center, Jichi Medical University, Saitama, Japan), which is a graphical user interface for R (The R Foundation for Statistical Computing, Vienna, Austria) [14]. For comparisons of two groups, Welch’s t test was performed. A value of p


